In a groundbreaking study that may pave new avenues for colorectal cancer treatment, researchers have unveiled a complex regulatory pathway involving the transcription factor RUNX1 and its critical role in modulating the tumor microenvironment. According to a recent publication in Cell Death Discovery, the investigation led by He, W., Zheng, L., Huang, W., and their colleagues presents compelling evidence that RUNX1 functions as a key suppressor of the STAT1-GITRL signaling axis, which orchestrates immune responses within colorectal carcinoma (CRC). This discovery sheds new light on the molecular crosstalk that governs immune evasion, a notorious hallmark of malignancy, particularly in CRC.
Colorectal cancer remains one of the most prevalent and lethal malignancies worldwide, partly due to its adeptness at subverting host immune defenses. While immunotherapy has revolutionized treatment for various cancer types, its efficacy in CRC has been notably limited. This limitation underscores the urgent need to understand the underlying immune landscape and identify molecular brakes imposed on antitumor immunity. The study focuses on RUNX1, a transcription factor previously recognized for its roles in hematopoiesis and leukemogenesis but less understood in solid tumors regarding immune regulation.
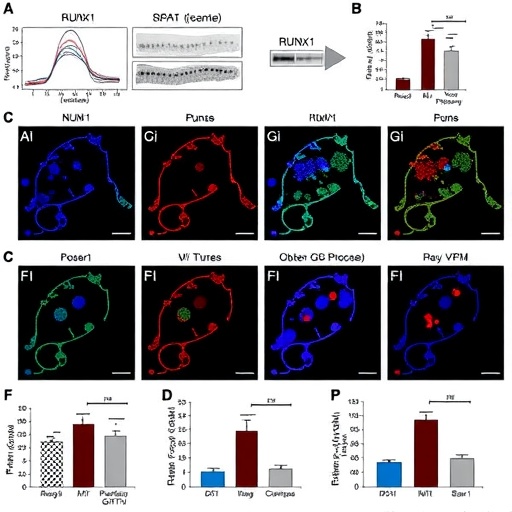
Through a series of integrative in vitro and in vivo experiments, the researchers demonstrated that RUNX1 expression is inversely correlated with the activation of the STAT1-GITRL signaling pathway in CRC tumor cells. STAT1, known primarily for propagating interferon-mediated responses, here emerges as a double-edged sword. While traditionally associated with immune activation, aberrant STAT1 activity, when coupled with the expression of Glucocorticoid-Induced TNFR-Related protein Ligand (GITRL), contributes to creating an immunosuppressive niche. This niche shields tumor cells from immune-mediated attack by promoting regulatory T cell (Treg) expansion and exhaustion of cytotoxic T lymphocytes.
Mechanistically, the team revealed that RUNX1 acts as a transcriptional repressor of STAT1, thereby reducing GITRL expression on tumor cells. Reduced GITRL limits the engagement of GITR receptors on effector immune cells, which in turn diminishes Treg amplification and preserves cytotoxic T cell function. Loss of RUNX1 in CRC cells unleashed the STAT1-GITRL axis, fostering an environment conducive to tumor immune escape. These findings highlight the dual role of STAT1 in cancer immunity and how transcriptional control via RUNX1 fine-tunes this balance.
Furthermore, the study employed patient-derived tumor samples and single-cell RNA sequencing to validate their observations in the clinical context. Analysis of CRC specimens confirmed that RUNX1 low tumors exhibited heightened STAT1 and GITRL expression, correlating with increased Treg infiltration and poorer prognostic outcomes. This translational insight lends significant weight to RUNX1’s potential as a biomarker for immune landscape characterization and therapeutic targeting.
To explore therapeutic implications, the researchers manipulated RUNX1 expression experimentally and observed that restoring RUNX1 function reactivated antitumor immunity and suppressed tumor growth in murine CRC models. Importantly, these effects were abrogated when the STAT1-GITRL axis was simultaneously inhibited, underscoring the pathway’s centrality in the immunomodulatory effects mediated by RUNX1. This positions RUNX1 as a promising candidate for strategies aimed at reinvigorating immune surveillance.
These discoveries challenge the previously simplistic view of STAT1 as merely a facilitator of antitumor effects and reveal a nuanced, context-dependent role. By delineating how RUNX1 restrains STAT1-induced immunosuppressive signaling, the study opens new perspectives on how tumor intrinsic factors shape immune microenvironments. It also suggests that therapeutic interventions could be tailored to modify this transcriptional control nexus, enhancing responsiveness to immunotherapy.
Importantly, the work delves into the epigenetic landscape dictating RUNX1 expression, identifying that hypermethylation of RUNX1 promoter regions in CRC contributes to its downregulation. This epigenetic silencing corroborates with diminished tumor immune surveillance and supports the rationale for combining epigenetic modifiers with immunotherapeutic regimens to restore RUNX1 function and immune competence.
The STAT1-GITRL pathway’s role extends beyond CRC, and the findings may have broader implications across other malignancies characterized by immune evasion. Future investigations are warranted to evaluate if RUNX1 modulation similarly impacts immune dynamics in diverse tumor types. Additionally, the prospect of developing small molecules or gene therapies aimed at boosting RUNX1 activity presents an exciting frontier.
Crucially, this study also highlights the intricate interplay between transcription factors and immune checkpoints within the tumor microenvironment. By focusing on the transcriptional circuits that govern ligand expression impacting immune cell phenotypes, the research adds a layer of complexity to our understanding of immune checkpoints, potentially guiding the design of novel checkpoint inhibitors or combinational approaches.
The implications extend into understanding resistance mechanisms to existing immunotherapies. Tumors harboring suppressed RUNX1 activity and an activated STAT1-GITRL circuit might inherently resist PD-1/PD-L1 blockade, explaining clinical variability. Patient stratification based on RUNX1 status could enhance personalized medicine approaches, optimizing treatment efficacy.
This study integrates molecular biology, immunology, and clinical oncology, underscoring the importance of interdisciplinary approaches in deciphering cancer biology. It exemplifies how basic science discoveries translate into actionable insights with tangible translational potential, offering hope in improving outcomes for CRC patients.
In conclusion, the identification of RUNX1 as a key regulator tempering the STAT1-GITRL axis introduces a paradigm shift in how we perceive immune regulation within colorectal cancer. By revealing the transcriptional checks that dictate immunosuppressive signaling pathways, this work lays the foundation for innovative therapies designed to dismantle tumor immune evasion. As science moves toward more precise and effective cancer immunotherapies, understanding and harnessing such molecular circuits will be pivotal.
Subject of Research: The modulation of colorectal cancer immune microenvironment by transcription factor RUNX1 and its impact on STAT1-GITRL signaling.
Article Title: RUNX1 restrains STAT1-GITRL signaling to shape an immunosuppressive CRC microenvironment.
Article References:
He, W., Zheng, L., Huang, W. et al. RUNX1 restrains STAT1-GITRL signaling to shape an immunosuppressive CRC microenvironment. Cell Death Discov. (2026). https://doi.org/10.1038/s41420-026-03053-7
Image Credits: AI Generated