In the expansive domain of eukaryotic life, protists stand out as a vast and diverse group whose evolutionary trajectories remain enigmatic despite their fundamental ecological roles. These microscopic organisms, which exclude animals, plants, and fungi, form the backbone of the eukaryotic tree of life, offering critical insights into cellular complexity and evolutionary history. However, their study has been notably hindered by the challenges associated with their cultivation and observation, constraining our understanding of their biological and phylogenetic intricacies.
A groundbreaking study conducted by researchers at the University of Tsukuba has now illuminated a previously obscured branch within the eukaryotic tree. By isolating and cultivating a protist from a marine environment in Palau, the team successfully identified a new species belonging to the genus Glissandra, which they have named Glissandra oviformis. This discovery is not only significant for taxonomy but also pivotal for deciphering the morphological and genetic traits that define major eukaryotic lineages.
The isolation of G. oviformis was accomplished through meticulous sampling of a marine lake and subsequent laboratory cultivation. Such efforts are vital because many protists are notoriously difficult to culture outside their natural habitats due to complex symbiotic relationships or specific environmental requirements. Overcoming these barriers, the researchers managed to stabilize a culture strain amenable to extensive morphological and molecular analyses, setting the stage for a comprehensive phylogenetic study.
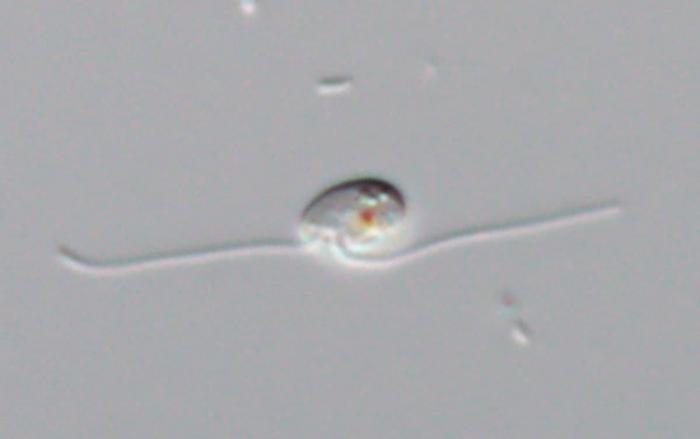
Utilizing high-resolution microscopy, including light and electron microscopic techniques, the team characterized the cellular architecture of G. oviformis. Notably, the cell membrane is lined with a distinctive sheet-like structure, a feature that had been hypothesized but rarely confirmed across related taxa. Additionally, the base of the organism’s flagella exhibited structural nuances consistent with members of the broader eukaryotic group known as CRuMs—a clade encompassing diverse and phylogenetically significant protists.
CRuMs represent one of the major eukaryotic clades but have remained poorly resolved due to the scant availability of cultured representatives and limited molecular data. The discovery of G. oviformis as a novel lineage within CRuMs thus addresses a critical gap in the tree of life. Molecular phylogenetic analyses based on a robust dataset of 340 protein-coding genes revealed that G. oviformis branches distinctly within CRuMs, underpinning the evolutionary divergence of this group from other eukaryotes.
This extensive phylogenomic approach employed concatenated protein sequences analyzed through advanced computational models, overcoming the confounding effects of gene paralogy and horizontal gene transfer that often obfuscate protist phylogenies. Such rigor provides unprecedented resolution and confidence in positioning G. oviformis within the eukaryotic framework, underscoring the organism’s pivotal role in elucidating character evolution at deep phylogenetic nodes.
Beyond phylogenetics, the morphological traits observed through electron microscopy furnish tangible links between G. oviformis and other CRuMs members. The shared presence of the membrane-adjacent sheet-like structure and specialized flagellar base morphology suggests conserved cellular features that may have been inherited from a common ancestor. These findings challenge previous assumptions that such structures arose convergently and emphasize the importance of ultrastructural studies in evolutionary biology.
The functional implications of these unique cellular architectures remain to be fully elucidated. Still, the configuration of the flagellar base and membrane structures likely plays a significant role in motility, environmental sensing, or feeding mechanisms. As G. oviformis is predatory, understanding these morphological adaptations can shed light on the ecological roles of CRuMs protists in marine ecosystems and their contributions to microbial food webs.
Research on G. oviformis also exemplifies the broader methodological advancements enabling protistology to re-enter the limelight of evolutionary studies. Large-scale molecular datasets combined with culturing of elusive species are unveiling the previously cryptic diversity within eukaryotic microbes. Such integrative approaches bridge morphological and molecular data, fostering a holistic framework to comprehend eukaryotic evolution and the complexities of cellular organization.
Significantly, this study highlights the rediscovery and comprehensive documentation of organisms with uncertain phylogenetic positions as a linchpin in reconstructing the early diversification of eukaryotes. By stabilizing cultures and thoroughly characterizing their cellular and genetic makeup, scientists can refine the evolutionary narrative, augmenting textbook models with empirical evidence from understudied microbial lineages.
In addition to broadening the taxonomic inventory, the insights gleaned from G. oviformis have potential downstream applications in biotechnology, ecology, and environmental monitoring. Protists are often sensitive bioindicators for ecosystem health, and understanding their evolutionary adaptations may inform conservation strategies, especially in sensitive marine environments such as the marine lakes of Palau.
The successful cultivation and analysis of G. oviformis thus represent a milestone not only for protist taxonomy but also for eukaryotic evolutionary biology at large. It exemplifies the power of combining fieldwork, microscopy, and molecular phylogenetics to uncover hidden biodiversity and unravel complex evolutionary patterns. As more novel protist lineages come to light, the evolutionary mosaic of eukaryotes will become increasingly resolved, revealing the origins of key cellular innovations and ecological strategies.
This pioneering work, spearheaded by the University of Tsukuba team, stands as a clarion call to the scientific community to intensify efforts toward eliciting hidden diversity within protists. As culturing techniques improve and sequencing becomes ever more precise, the enigmatic branches of the eukaryotic tree of life will progressively be illuminated, offering answers to long-standing questions about the evolution of cellular life forms.
The study of Glissandra oviformis epitomizes the intricate interplay of morphology and molecular data required to comprehend deeply rooted evolutionary relationships. It serves as a model case demonstrating that persistent cultivation and detailed characterization of protists can unlock new perspectives on evolutionary biology, reinforcing the critical role of microbial eukaryotes in the history of life on Earth.
Subject of Research: Protists, Eukaryotic Phylogenetics, CRuMs Clade
Article Title: Glissandra oviformis n. sp.: a novel predatory flagellate illuminates the character evolution within the eukaryotic clade CRuMs
News Publication Date: 4-Jun-2025
Web References:
https://doi.org/10.1098/rsob.250057
Image Credits: University of Tsukuba
Keywords: Protists, Molecular Phylogenetics, Taxonomies