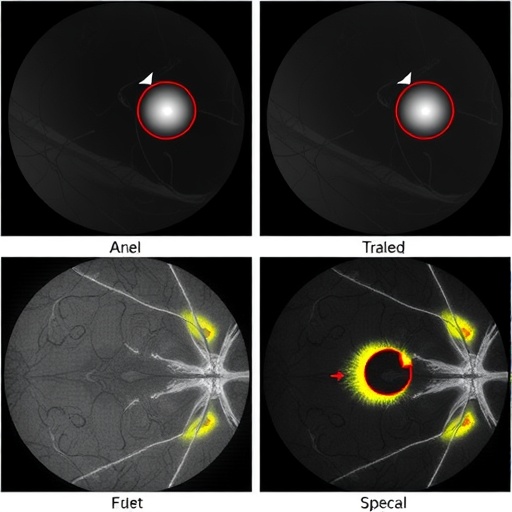
Automated lesion segmentation plays a crucial role in the early detection and management of diabetic retinopathy (DR), a leading cause of vision loss worldwide. However, current deep learning-based approaches often struggle with robustness, particularly in challenging cases where low contrast and imaging artifacts obscure lesion visibility. This vulnerability results in an increased rate of false positives, undermining the clinical utility of these systems. The root cause lies in the insufficient incorporation of anatomical knowledge, which is vital for discerning true pathological changes from noise and artifacts within retinal images.
To overcome this limitation, researchers have introduced a novel framework known as MedFuse, unveiled on March 15, 2026, in the prestigious journal Frontiers of Computer Science. The MedFuse system is designed to enhance the reliability of lesion segmentation by integrating explicit anatomical priors into the model’s decision-making process. Unlike traditional supervised methods that require painstakingly annotated training data, especially for intricate vascular structures, MedFuse adopts a zero-shot approach leveraging a multimodal large language model (LLM).
This zero-shot paradigm is groundbreaking because it eliminates the dependency on pixel-level vessel annotations, which are notoriously expensive and scarce due to the complexity of manual labeling. Instead, the LLM autonomously extracts precise vascular priors directly from raw retinal images. These priors function as stable anatomical anchors, providing a consistent structural reference that guides the segmentation network. By aligning the model’s visual feature extraction with these anatomical landmarks, MedFuse effectively differentiates true diabetic lesions from background noise, substantially reducing false positives.
The architecture of MedFuse is thoughtfully engineered to fuse multi-source data streams within a unified framework. It synergizes visual information with anatomical context, fostering a robust representation of retinal pathology. The fusion process leverages the power of large language models to interpret subtle vascular patterns, which are otherwise challenging to capture through conventional convolutional neural networks alone. This fusion enables a mechanism-guided segmentation approach that transcends the limitations of purely data-driven models.
Experimental validation of MedFuse was conducted on two well-established datasets: DDR (Diabetic Retinopathy Detection) and IDRID (Indian Diabetic Retinopathy Image Dataset). These datasets are benchmarks in the field, known for their diversity and complexity. The results demonstrated that MedFuse achieved significant improvements in segmentation accuracy and robustness compared to leading baseline methods. Notably, the alignment of visual features with vascular priors led to more faithful lesion delineation, particularly in regions marred by low contrast or artifacts.
The enhanced performance of MedFuse is not merely about accuracy but also data efficiency. By leveraging anatomical priors, the framework reduces the need for large volumes of annotated training data, which is a significant bottleneck in medical imaging AI development. This efficiency accelerates the pathway toward clinical deployment, offering a scalable solution adaptable to diverse patient populations and imaging conditions.
Moreover, MedFuse’s mechanism-guided design holds promise for interpretability, a critical factor in medical applications. By explicitly modeling anatomical structures, the system offers clinicians intuitive insights into its decision rationale. This transparency fosters trust and facilitates integration into clinical workflows, where clear explanations of AI predictions are mandatory for regulatory approval and user acceptance.
The implications of this research extend beyond diabetic retinopathy. The conceptual framework of fusing multimodal data with anatomical knowledge through large language models can be generalized to other medical imaging tasks. Diseases that present with complex anatomical variations or subtle pathology could benefit from such mechanism-guided approaches, potentially catalyzing a new era of robust, explainable AI in healthcare.
The research team behind MedFuse comprises experts in computer vision, medical imaging, and artificial intelligence. Their multidisciplinary collaboration underscores the importance of cross-domain knowledge for tackling challenging problems in biomedical data analysis. The publication in Frontiers of Computer Science, a journal celebrated for cutting-edge innovations, highlights the significance and forward-looking nature of this work.
This advancement arrives at a pivotal moment when the medical community increasingly harnesses artificial intelligence to augment diagnostic accuracy and efficiency. MedFuse sets a benchmark for future research by demonstrating that incorporating explicit anatomical priors, intelligently generated by state-of-the-art language models, can bridge the gap between high performance and clinical reliability.
In conclusion, MedFuse represents a pioneering step toward building intelligent, anatomically aware medical imaging systems for diabetic retinopathy lesion segmentation. Its innovative use of zero-shot multimodal LLMs to generate vascular priors marks a paradigm shift from data-heavy supervision to knowledge-infused learning. As this framework matures, it promises to facilitate earlier diagnosis, personalized disease monitoring, and ultimately improved patient outcomes in diabetic eye care.
Subject of Research: Not applicable
Article Title: MedFuse: a multi-source data fusion framework for diabetic retinopathy lesion segmentation
News Publication Date: 15-Mar-2026
Web References:
10.1007/s11704-025-51690-5
Image Credits: HIGHER EDUCATION PRESS
Keywords: Computer science, diabetic retinopathy, lesion segmentation, deep learning, anatomical priors, large language model, multimodal fusion, zero-shot learning, medical imaging AI