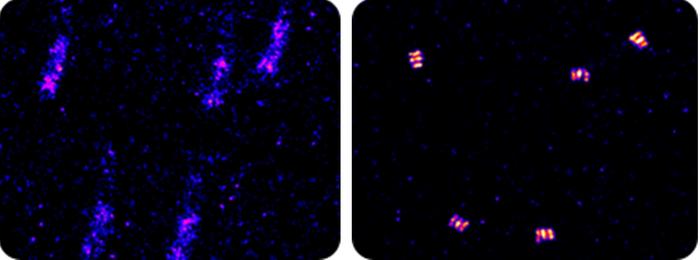
When trying to measure molecular structures with nanometer precision, every bit of noise shows up in the data: someone walking past the microscope, tiny vibrations in the building and even the traffic outside. A new processing technique removes noise from optical microscope data in real time, allowing scientists to track individual molecules over 10 times more precisely than was possible before.

Credit: The Grainger College of Engineering at the University of Illinois Urbana-Champaign
When trying to measure molecular structures with nanometer precision, every bit of noise shows up in the data: someone walking past the microscope, tiny vibrations in the building and even the traffic outside. A new processing technique removes noise from optical microscope data in real time, allowing scientists to track individual molecules over 10 times more precisely than was possible before.
A team of bioengineering researchers at the University of Illinois Urbana-Champaign has introduced an algorithm called adaptive intersection maximization, or AIM, that removes high-frequency noise from super-resolution optical microscope data much faster than standard methods and results in much higher image resolutions. The algorithm will enable scientists to study chemical and biological systems far more easily and precisely than was possible before. This research was published in the journal Science Advances.
“At first, we just wanted to develop a fast algorithm because our lab produces too much data for traditional algorithms to handle, but we found that AIM can also achieve sub-nanometer precision, which is unheard of in our field,” said Hongqiang Ma, a research professor of bioengineering and the study’s lead author. “In addition, it doesn’t require immense computing power like traditional tools. It can run on a laptop. We want to make this a plug-and-play tool for all microscope users.”
In recent decades, the single-molecule localization microscopy technique has enabled scientists to visualize molecular-scale structures, surpassing what was thought to be a fundamental limitation of optical microscopes. However, it is limited in practice by uncontrollable noise, or “drift,” that essentially blurs the images and prevents super-resolution microscopy from reaching its highest resolution.
“Single-molecule localization actually uses a fairly simple instrument, but the tricky part that really impacts image resolution is drift,” said Yang Liu, a bioengineering professor and the project lead. “Many researchers only remove low-frequency drift. Removing the high-frequency drift – minute vibrations caused by environmental noise – is computationally intensive and requires large amounts of time and resources.”
Standard methods for removing drift are based on the mathematical correlations between image frames. According to Liu, the microscopes in her laboratory generate such a large volume of image data that image correlation methods take days even with supercomputing resources.
AIM also compares adjacent frames, but it proceeds by putting each data point at the center of a circle (defined by localization precision) and looking for points inside that circle in other frames. Overlapping points within the “radius of intersection” are condensed into a single localization. Then, the process is repeated once more with the condensed points. These steps use minimal computational resources, and they are faster than the acquisition time of a microscope camera. So, drift-corrected images can be produced in what is effectively real time.
The researchers tested AIM using both simulated data and structures called DNA origami that have well-defined features. The algorithm successfully localized the structures, and the degree of precision, less than 1 nanometer, turned out to be much higher than standard image correlation methods, about 10 nanometers.
Liu’s laboratory will incorporate AIM into high-throughput microscopy techniques being developed for enhanced disease detection. However, Liu also believes that the algorithm will find uses throughout biology and bioengineering. “It’s a fast and easy-to-use tool, and we want to make it widely accessible for the entire community,” she said. “We are making our software publicly accessible. We want people to get the boost in their image resolution just from this one bit of post-processing.”
The study, “Towards drift-free high-throughput nanoscopy through adaptive intersection maximization,” is available online. DOI: 10.1126/sciadv.adm7765
Maomao Chen of the University of Pittsburgh and Phuong Nguyen of Illinois also contributed to this work.
Liu is also affiliated with the department of electrical & computer engineering and the Cancer Center at Illinois. Ma and Nguyen are also affiliated with the Beckman Institute for Advanced Science and Technology at Illinois.
Support was provided by the National Institutes of Health.
Journal
Science Advances
Article Title
Toward drift-free high-throughput nanoscopy through adaptive intersection maximization
Article Publication Date
24-May-2024