In the rapidly evolving field of computational biology, the challenge of predicting antimicrobial resistance in pathogens has gained critical importance, especially with the increasing prevalence of resistant strains of organisms such as Salmonella. A groundbreaking study published in the journal Engineering introduces an innovative predictive platform that combines the formidable capabilities of large language models (LLMs) with the advancements of quantum computing. This research, spearheaded by a team from Sichuan University under the leadership of Le Zhang, represents a significant leap forward in addressing one of the most pressing public health concerns of our time.
The rise of antimicrobial resistance, particularly in common foodborne pathogens like Salmonella, has been exacerbated by the overuse of antibiotics in agriculture and human medicine. The genetic mutations that accompany such practices have led to strains that are resistant to standard treatment protocols. Traditionally, predicting this resistance has relied on bacterial antimicrobial susceptibility tests (ASTs), which can be resource-intensive and time-consuming. Furthermore, current genomic analyses often fall prey to issues of overfitting, largely due to the high-dimensional nature of whole-genome sequencing (WGS) data.
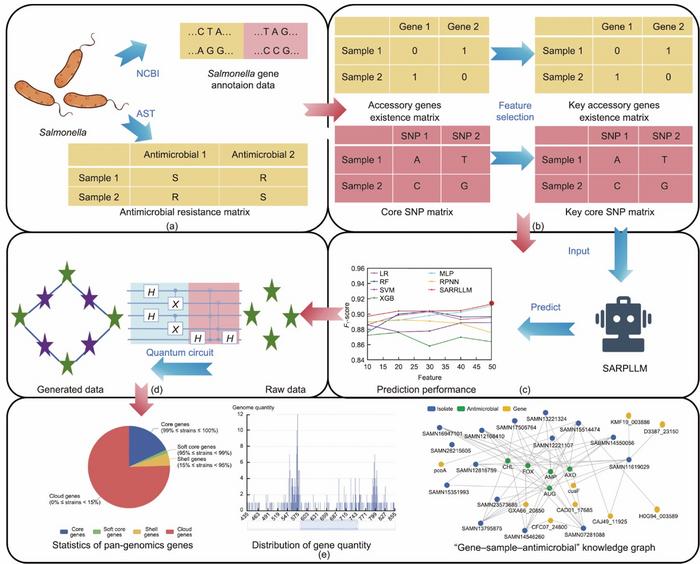
Given these complexities, the researchers turned to a dual-layered feature-selection process, which is crucial for extracting pivotal genetic information relevant to antimicrobial resistance. The initial step involves the application of a chi-square test to sift through vast datasets and identify the key resistance genes within the Salmonella genome. This technique is further complemented by a conditional mutual information maximization approach that enhances the process of identifying critical features.
Building on the initial findings, the team then developed a sophisticated algorithm, dubbed the Salmonella Antimicrobial Resistance Predictive Large Language Model (SARPLLM). This model is rooted in the Qwen2 LLM, augmented with low-rank adaptation (LoRA) technology, allowing for the conversion of complex genomic data into a sentence-like structure that the model can effectively analyze. By translating genetic variations and resistance markers into this textual format, the SARPLLM can predict antimicrobial resistance with a level of accuracy that far surpasses prevailing methodologies.
However, despite the promise of this innovative approach, the study does not shy away from acknowledging inherent challenges. Notably, the datasets used in genomic analyses often exhibit an imbalance in the representation of antimicrobial-resistant versus sensitive strains of Salmonella. This discrepancy can severely limit the efficacy of predictive models. To tackle this issue, the researchers developed the QSMOTEN algorithm, an adaptation of the existing SMOTEN algorithm that utilizes the principles of quantum computing. This novel algorithm encodes sample features into quantum states, enabling efficient computation of distances between samples. By drastically reducing the complexity of these calculations from a linear to a logarithmic scale, QSMOTEN effectively enhances the processing of complex high-dimensional data associated with WGS.
The remarkable findings presented in this study also encompass the creation of a user-oriented online platform designed for antimicrobial resistance prediction. Utilizing the Django framework for backend operations and Echarts for robust knowledge graph visualization, the platform features multiple modules. These include an intuitive predictive module for antimicrobial resistance, a dedicated space for presenting pan-genomics analysis results, an interactive gene-sample-antimicrobial knowledge-graph module, and a streamlined data download feature. This user-friendly interface enables researchers and practitioners to upload gene feature files for real-time predictions, bolstered by extensive data visualization capabilities.
Impressive experimental results underpin the claims made by the researchers regarding the superior performance of the SARPLLM algorithm in predicting antimicrobial resistance. With elevated F1-scores across various antimicrobial drugs, it is evident that the integration of LLMs into this domain can yield significant advancements. Moreover, the efficiency of the QSMOTEN algorithm in accurately measuring sample similarities has been validated on both virtual and physical quantum computing platforms, demonstrating the transformative potential of this technology in the realm of biological data analysis and augmentation.
Nonetheless, the authors are candid in recognizing the limitations that currently exist within their work. The intricate biological and genetic landscapes surrounding antimicrobial resistance present considerable challenges, particularly concerning the limitations of LLMs in fully grasping complex domain knowledge. The efficacy of these models is tightly intertwined with the quality and scope of the training data employed, further emphasizing the necessity for continuous improvements in this field of study. The nascent stage of quantum computing technology also remains a noteworthy consideration, underscoring the need for ongoing research aimed at refining integration processes and establishing more robust quantum infrastructures.
As the healthcare landscape grapples with the rising tide of antimicrobial resistance, this study represents a monumental shift towards more predictive and proactive healthcare solutions. The implications of leveraging advanced computational power to preemptively identify strains with resistance potential could not only pave the way for more effective treatment strategies but also inform public health policies aimed at reducing the prevalence of resistant pathogens in the environment.
Moving forward, the researchers plan to concentrate their efforts on enhancing the predictive platform by integrating multi-source datasets alongside domain-specific knowledge. This approach may significantly bolster the accuracy of resistance predictions, further mitigating the healthcare crisis posed by antimicrobial-resistant strains. Additionally, advancements in quantum hardware will be pivotal in achieving more streamlined and stable performance for such complex applications.
The innovative strides made in this research signify a promising trajectory towards harnessing cutting-edge technologies like quantum computing and LLMs to tackle some of the most formidable challenges in public health. The potential of these methods extends beyond the current study, setting a precedent for future explorations into the intersection of computational techniques and biological insights, ultimately aiming for better health outcomes globally.
—
Subject of Research: Prediction of Salmonella Antimicrobial Resistance
Article Title: Developing a Predictive Platform for Salmonella Antimicrobial Resistance Based on a Large Language Model and Quantum Computing
News Publication Date: 28-Jan-2025
Web References: https://doi.org/10.1016/j.eng.2025.01.013
References: 10.1016/j.eng.2025.01.013
Image Credits: Yujie You et al.
Keywords: Salmonella, antimicrobial resistance, large language models, quantum computing, predictive algorithms, pan-genomics, QSMOTEN, SARPLLM.